|
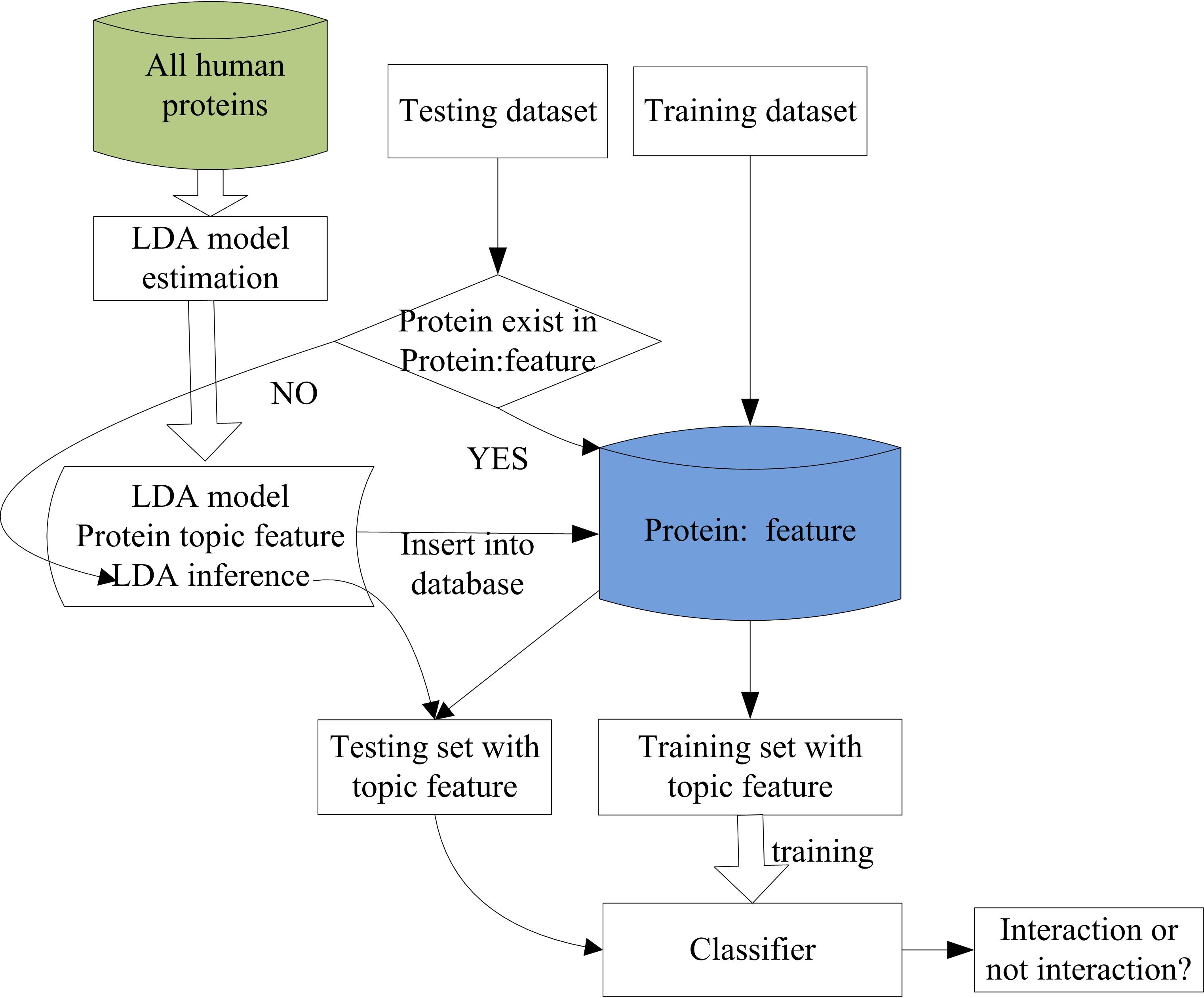
Protein-protein interaction (PPI) is believed at the core of entire interactomic system of any living organism. Although there are many human protein-protein interaction links have been experimentally determined, the number is still relatively very few compared to the estimation that there will be ~300,000 protein-protein interactions in human beings. Hence, it is still an urgent and challenge work to develop automated computational methods to accurately and efficiently predict protein-protein interactions. In this paper, we propose a novel hierarchical LDA-RF (latent dirichlet allocation-random forest) model to predict human protein-protein interactions from protein primary sequences directly, which is featured by high success rate and strong ability for handling large-scale datasets by digging the hidden internal structures buried into the noisy amino acid sequences in the low dimension latent semantic space. Firstly, the local sequential features represented by conjoint triads are constructed from sequences. Then the generative LDA model is used to project the original feature space into the latent semantic space to obtain low dimensional latent topic features, which reflect the hidden structures of the protein. Finally, the powerful random forest model is used to predict the probability for interaction of two proteins based on the concatenation of the two separate latent topic features. Our results show that the proposed latent topic feature is very promising for PPI prediction and could also become a powerful strategy to deal with many other bioinformatics problems.
|